Which Stage Is Most Associated With a Cell That Is Unable to Divide Again Such as a Nerve Cell


Onion (Allium) cells in different phases of the cell cycle. Growth in an 'organism' is advisedly controlled past regulating the cell bike.
Prison cell bike in Deinococcus radiodurans
The jail cell cycle, or cell-division cycle, is the series of events that take identify in a jail cell that cause it to dissever into two daughter cells. These events include the duplication of its Deoxyribonucleic acid (DNA replication) and some of its organelles, and subsequently the partitioning of its cytoplasm and other components into two girl cells in a process chosen cell division.
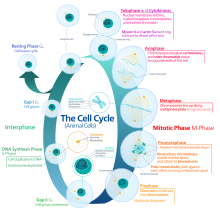
In cells with nuclei (eukaryotes, i.e., animal, plant, fungal, and protist cells), the cell cycle is divided into two chief stages: interphase and the mitotic (M) phase (including mitosis and cytokinesis). During interphase, the cell grows, accumulating nutrients needed for mitosis, and replicates its Deoxyribonucleic acid and some of its organelles. During the mitotic phase, the replicated chromosomes, organelles, and cytoplasm separate into 2 new daughter cells. To ensure the proper replication of cellular components and partitioning, there are control mechanisms known as prison cell bicycle checkpoints after each of the primal steps of the bike that determine if the jail cell can progress to the next phase.
In cells without nuclei (prokaryotes, i.eastward., bacteria and archaea), the jail cell cycle is divided into the B, C, and D periods. The B flow extends from the finish of cell division to the beginning of DNA replication. DNA replication occurs during the C period. The D period refers to the stage between the cease of Deoxyribonucleic acid replication and the splitting of the bacterial cell into ii daughter cells.[ane]
The prison cell-division bicycle is a vital process by which a single-celled fertilized egg develops into a mature organism, likewise as the process past which pilus, skin, blood cells, and some internal organs are regenerated and healed (with possible exception of nerves; see nerve damage). After cell partitioning, each of the daughter cells begin the interphase of a new cell cycle. Although the various stages of interphase are not unremarkably morphologically distinguishable, each stage of the cell cycle has a singled-out set up of specialized biochemical processes that prepare the cell for initiation of the cell sectionalisation.
Phases [edit]
The eukaryotic cell bicycle consists of 4 distinct phases: Gi phase, S stage (synthesis), Thou2 phase (collectively known every bit interphase) and M phase (mitosis and cytokinesis). Thousand phase is itself composed of two tightly coupled processes: mitosis, in which the cell'due south nucleus divides, and cytokinesis, in which the cell'southward cytoplasm divides forming ii daughter cells. Activation of each phase is dependent on the proper progression and completion of the previous i. Cells that have temporarily or reversibly stopped dividing are said to have entered a country of quiescence called Grand0 phase.

| State | Phase | Abbreviation | Description |
|---|---|---|---|
| Resting | Gap 0 | M0 | A phase where the jail cell has left the bicycle and has stopped dividing. |
| Interphase | Gap i | Gone | Cells increment in size in Gap ane. The G1 checkpoint control mechanism ensures that everything is ready for Dna synthesis. |
| Synthesis | S | Deoxyribonucleic acid replication occurs during this phase. | |
| Gap 2 | 10002 | During the gap between Dna synthesis and mitosis, the prison cell will continue to abound. The Thouii checkpoint control machinery ensures that everything is fix to enter the M (mitosis) phase and dissever. | |
| Cell division | Mitosis | M | Jail cell growth stops at this stage and cellular energy is focused on the orderly partitioning into 2 daughter cells. A checkpoint in the middle of mitosis (Metaphase Checkpoint) ensures that the cell is ready to consummate cell division. |
After cell sectionalization, each of the daughter cells brainstorm the interphase of a new cycle. Although the various stages of interphase are not usually morphologically distinguishable, each phase of the cell cycle has a singled-out set of specialized biochemical processes that fix the cell for initiation of cell division.
G0 phase (quiescence) [edit]

M0 is a resting phase where the cell has left the cycle and has stopped dividing. The cell bike starts with this phase. Not-proliferative (non-dividing) cells in multicellular eukaryotes mostly enter the quiescent Chiliad0 state from One thousand1 and may remain quiescent for long periods of time, possibly indefinitely (as is often the case for neurons). This is very common for cells that are fully differentiated. Some cells enter the G0 stage semi-permanently and are considered postal service-mitotic, e.one thousand., some liver, kidney, and stomach cells. Many cells practise not enter M0 and continue to carve up throughout an organism's life, due east.g., epithelial cells.
The word "mail-mitotic" is sometimes used to refer to both quiescent and senescent cells. Cellular senescence occurs in response to Dna damage and external stress and ordinarily constitutes an arrest in Gi. Cellular senescence may make a cell'due south progeny nonviable; it is often a biochemical alternative to the cocky-destruction of such a damaged cell past apoptosis.
Interphase [edit]
Interphase stand for the phase between two successive M phases. Interphase is a series of changes that takes place in a newly formed cell and its nucleus before it becomes capable of partition over again. Information technology is also chosen preparatory stage or intermitosis. Typically interphase lasts for at least 91% of the total fourth dimension required for the cell cycle.
Interphase proceeds in 3 stages, G1, S, and Grand2, followed by the bicycle of mitosis and cytokinesis. The cell's nuclear Dna contents are duplicated during S stage.
Mane phase (Showtime growth stage or Mail service mitotic gap phase) [edit]
The first phase inside interphase, from the end of the previous M stage until the offset of DNA synthesis, is called Gone (G indicating gap). Information technology is also called the growth stage. During this phase, the biosynthetic activities of the cell, which are considerably slowed downward during M stage, resume at a high rate. The duration of Kane is highly variable, even amongst different cells of the aforementioned species.[three] In this stage, the cell increases its supply of proteins, increases the number of organelles (such as mitochondria, ribosomes), and grows in size. In Chiliad1 phase, a cell has three options.
- To go along cell cycle and enter Southward phase
- Stop cell wheel and enter Chiliad0 stage for undergoing differentiation.
- Go arrested in Gi phase hence it may enter G0 stage or re-enter cell wheel.
The deciding betoken is called check bespeak (Restriction signal). This cheque point is chosen the restriction point or Outset and is regulated by G1/S cyclins, which cause transition from G1 to S stage. Passage through the Gane check point commits the jail cell to division.
S stage (DNA replication) [edit]
The ensuing S stage starts when DNA synthesis commences; when it is complete, all of the chromosomes have been replicated, i.e., each chromosome consists of two sis chromatids. Thus, during this stage, the amount of Dna in the prison cell has doubled, though the ploidy and number of chromosomes are unchanged. Rates of RNA transcription and poly peptide synthesis are very low during this phase. An exception to this is histone production, most of which occurs during the S phase.[iv] [v] [six]
Thousand2 phase (growth) [edit]
G2 phase occurs after DNA replication and is a catamenia of protein synthesis and rapid prison cell growth to set up the cell for mitosis. During this phase microtubules begin to reorganize to form a spindle (preprophase). Before proceeding to mitotic stage, cells must be checked at the Gii checkpoint for any DNA damage inside the chromosomes. The G2 checkpoint is mainly regulated past the tumor poly peptide p53. If the DNA is damaged, p53 will either repair the Deoxyribonucleic acid or trigger the apoptosis of the cell. If p53 is dysfunctional or mutated, cells with damaged DNA may continue through the cell cycle, leading to the evolution of cancer.
Mitotic phase (chromosome separation) [edit]
The relatively brief M phase consists of nuclear division (karyokinesis). It is a relatively short period of the cell bike. Thou phase is complex and highly regulated. The sequence of events is divided into phases, corresponding to the completion of ane set of activities and the offset of the side by side. These phases are sequentially known equally:
- prophase
- prometaphase
- metaphase
- anaphase
- telophase
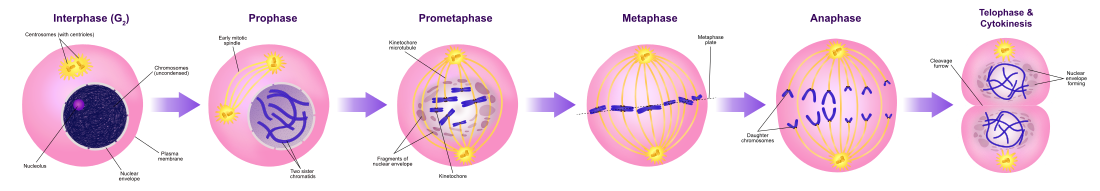
Mitosis is the process by which a eukaryotic cell separates the chromosomes in its prison cell nucleus into two identical sets in two nuclei.[7] During the process of mitosis the pairs of chromosomes condense and attach to microtubules that pull the sister chromatids to contrary sides of the prison cell.[8]
Mitosis occurs exclusively in eukaryotic cells, but occurs in different ways in different species. For example, fauna cells undergo an "open up" mitosis, where the nuclear envelope breaks downwards before the chromosomes separate, while fungi such as Aspergillus nidulans and Saccharomyces cerevisiae (yeast) undergo a "closed" mitosis, where chromosomes divide inside an intact cell nucleus.[nine]
Cytokinesis phase (separation of all cell components) [edit]
Mitosis is immediately followed past cytokinesis, which divides the nuclei, cytoplasm, organelles and cell membrane into 2 cells containing roughly equal shares of these cellular components. Mitosis and cytokinesis together define the division of the mother jail cell into two daughter cells, genetically identical to each other and to their parent cell. This accounts for approximately 10% of the cell bike.
Considering cytokinesis usually occurs in conjunction with mitosis, "mitosis" is often used interchangeably with "K phase". Nevertheless, there are many cells where mitosis and cytokinesis occur separately, forming single cells with multiple nuclei in a process called endoreplication. This occurs most notably amid the fungi and slime molds, but is found in diverse groups. Even in animals, cytokinesis and mitosis may occur independently, for example during certain stages of fruit fly embryonic development.[x] Errors in mitosis can result in cell death through apoptosis or cause mutations that may lead to cancer.
Regulation of eukaryotic cell cycle [edit]

Levels of the three major cyclin types oscillate during the cell cycle (superlative), providing the basis for oscillations in the cyclin–Cdk complexes that drive prison cell-wheel events (lesser). In full general, Cdk levels are constant and in large excess over cyclin levels; thus, cyclin–Cdk complexes class in parallel with cyclin levels. The enzymatic activities of cyclin–Cdk complexes besides tend to rise and autumn in parallel with cyclin levels, although in some cases Cdk inhibitor proteins or phosphorylation introduce a delay between the germination and activation of cyclin–Cdk complexes. Formation of active G1/Due south–Cdk complexes commits the cell to a new division cycle at the Start checkpoint in belatedly G1. G1/S–Cdks then activate the S–Cdk complexes that initiate DNA replication at the beginning of S stage. M–Cdk activation occurs afterwards the completion of S phase, resulting in progression through the G2/Chiliad checkpoint and assembly of the mitotic spindle. APC activation then triggers sister-chromatid separation at the metaphase-to-anaphase transition. APC activity also causes the destruction of S and K cyclins and thus the inactivation of Cdks, which promotes the completion of mitosis and cytokinesis. APC activeness is maintained in G1 until G1/S–Cdk activity rises again and commits the cell to the next cycle. This scheme serves but as a general guide and does not apply to all prison cell types.
Regulation of the cell cycle involves processes crucial to the survival of a cell, including the detection and repair of genetic impairment besides as the prevention of uncontrolled jail cell division. The molecular events that control the cell cycle are ordered and directional; that is, each process occurs in a sequential way and it is incommunicable to "reverse" the cycle.
Role of cyclins and CDKs [edit]
Two key classes of regulatory molecules, cyclins and cyclin-dependent kinases (CDKs), determine a jail cell's progress through the prison cell bike.[11] Leland H. Hartwell, R. Timothy Chase, and Paul M. Nurse won the 2001 Nobel Prize in Physiology or Medicine for their discovery of these fundamental molecules.[12] Many of the genes encoding cyclins and CDKs are conserved among all eukaryotes, but in full general, more complex organisms have more than elaborate cell cycle command systems that incorporate more individual components. Many of the relevant genes were beginning identified by studying yeast, particularly Saccharomyces cerevisiae;[xiii] genetic nomenclature in yeast dubs many of these genes cdc (for "prison cell division cycle") followed by an identifying number, e.g. cdc25 or cdc20.
Cyclins form the regulatory subunits and CDKs the catalytic subunits of an activated heterodimer; cyclins take no catalytic activity and CDKs are inactive in the absence of a partner cyclin. When activated by a bound cyclin, CDKs perform a common biochemical reaction called phosphorylation that activates or inactivates target proteins to orchestrate coordinated entry into the side by side phase of the prison cell cycle. Different cyclin-CDK combinations determine the downstream proteins targeted. CDKs are constitutively expressed in cells whereas cyclins are synthesised at specific stages of the cell cycle, in response to various molecular signals.[14]
General mechanism of cyclin-CDK interaction [edit]
Upon receiving a pro-mitotic extracellular point, K1 cyclin-CDK complexes become active to fix the jail cell for S stage, promoting the expression of transcription factors that in plough promote the expression of South cyclins and of enzymes required for Deoxyribonucleic acid replication. The Grand1 cyclin-CDK complexes likewise promote the degradation of molecules that function as S stage inhibitors by targeting them for ubiquitination. Once a protein has been ubiquitinated, information technology is targeted for proteolytic degradation by the proteasome. However, results from a recent study of E2F transcriptional dynamics at the single-cell level argue that the role of G1 cyclin-CDK activities, in particular cyclin D-CDK4/6, is to tune the timing rather than the commitment of cell cycle entry.[fifteen]
Active Southward cyclin-CDK complexes phosphorylate proteins that brand upwardly the pre-replication complexes assembled during Gi phase on DNA replication origins. The phosphorylation serves two purposes: to activate each already-assembled pre-replication complex, and to prevent new complexes from forming. This ensures that every portion of the cell's genome will be replicated once and merely once. The reason for prevention of gaps in replication is fairly articulate, because daughter cells that are missing all or role of crucial genes will die. Nevertheless, for reasons related to factor re-create number effects, possession of extra copies of sure genes is also deleterious to the daughter cells.
Mitotic cyclin-CDK complexes, which are synthesized simply inactivated during S and G2 phases, promote the initiation of mitosis past stimulating downstream proteins involved in chromosome condensation and mitotic spindle assembly. A critical complex activated during this process is a ubiquitin ligase known as the anaphase-promoting complex (APC), which promotes degradation of structural proteins associated with the chromosomal kinetochore. APC also targets the mitotic cyclins for degradation, ensuring that telophase and cytokinesis tin can go along.[sixteen]
Specific activity of cyclin-CDK complexes [edit]
Cyclin D is the first cyclin produced in the cells that enter the prison cell cycle, in response to extracellular signals (e.chiliad. growth factors). Cyclin D levels stay low in resting cells that are non proliferating. Additionally, CDK4/6 and CDK2 are as well inactive because CDK4/6 are bound by INK4 family members (east.g., p16), limiting kinase activity. Meanwhile, CDK2 complexes are inhibited by the CIP/KIP proteins such every bit p21 and p27,[17] When it is time for a prison cell to enter the cell cycle, which is triggered by a mitogenic stimuli, levels of cyclin D increase. In response to this trigger, cyclin D binds to existing CDK4/6, forming the active cyclin D-CDK4/vi complex. Cyclin D-CDK4/vi complexes in turn mono-phosphorylates the retinoblastoma susceptibility protein (Rb) to pRb. The un-phosphorylated Rb neoplasm suppressor functions in inducing cell bike leave and maintaining G0 abort (senescence).[18]
In the terminal few decades, a model has been widely accustomed whereby pRB proteins are inactivated by cyclin D-Cdk4/6-mediated phosphorylation. Rb has fourteen+ potential phosphorylation sites. Cyclin D-Cdk 4/6 progressively phosphorylates Rb to hyperphosphorylated land, which triggers dissociation of pRB–E2F complexes, thereby inducing G1/S cell cycle gene expression and progression into S phase.[19]
However, scientific observations from a recent written report testify that Rb is present in three types of isoforms: (1) un-phosphorylated Rb in G0 state; (2) mono-phosphorylated Rb, too referred to as "hypo-phosphorylated' or 'partially' phosphorylated Rb in early G1 state; and (3) inactive hyper-phosphorylated Rb in late G1 country.[20] [21] [22] In early G1 cells, mono-phosphorylated Rb exits every bit xiv different isoforms, ane of each has distinct E2F binding affinity.[22] Rb has been found to associate with hundreds of different proteins[23] and the thought that unlike mono-phosphorylated Rb isoforms have dissimilar protein partners was very appealing.[24] A recent report confirmed that mono-phosphorylation controls Rb's association with other proteins and generates functional distinct forms of Rb.[25] All dissimilar mono-phosphorylated Rb isoforms inhibit E2F transcriptional program and are able to abort cells in G1-phase. Importantly, unlike mono-phosphorylated forms of RB take distinct transcriptional outputs that are extended beyond E2F regulation.[25]
In general, the binding of pRb to E2F inhibits the E2F target gene expression of certain G1/Southward and S transition genes including E-type cyclins. The partial phosphorylation of RB de-represses the Rb-mediated suppression of E2F target gene expression, begins the expression of cyclin E. The molecular mechanism that causes the cell switched to cyclin E activation is currently not known, only as cyclin E levels rise, the active cyclin Due east-CDK2 circuitous is formed, bringing Rb to be inactivated past hyper-phosphorylation.[22] Hyperphosphorylated Rb is completely dissociated from E2F, enabling farther expression of a broad range of E2F target genes are required for driving cells to continue into South phase [1]. Recently, it has been identified that cyclin D-Cdk4/6 binds to a C-final alpha-helix region of Rb that is just distinguishable to cyclin D rather than other cyclins, cyclin E, A and B.[26] This observation based on the structural assay of Rb phosphorylation supports that Rb is phosphorylated in a different level through multiple Cyclin-Cdk complexes. This also makes feasible the electric current model of a simultaneous switch-like inactivation of all mono-phosphorylated Rb isoforms through one type of Rb hyper-phosphorylation machinery. In addition, mutational analysis of the cyclin D- Cdk iv/6 specific Rb C-last helix shows that disruptions of cyclin D-Cdk 4/6 bounden to Rb prevents Rb phosphorylation, arrests cells in G1, and bolsters Rb'south functions in tumor suppressor.[26] This cyclin-Cdk driven cell cycle transitional mechanism governs a cell committed to the cell cycle that allows cell proliferation. A cancerous cell growth often accompanies with deregulation of Cyclin D-Cdk 4/6 activity.
The hyperphosphorylated Rb dissociates from the E2F/DP1/Rb complex (which was bound to the E2F responsive genes, effectively "blocking" them from transcription), activating E2F. Activation of E2F results in transcription of various genes like cyclin E, cyclin A, DNA polymerase, thymidine kinase, etc. Cyclin E thus produced binds to CDK2, forming the cyclin E-CDK2 circuitous, which pushes the jail cell from Thouane to Southward stage (10001/S, which initiates the Thousand2/1000 transition).[27] Cyclin B-cdk1 complex activation causes breakdown of nuclear envelope and initiation of prophase, and later, its deactivation causes the cell to exit mitosis.[14] A quantitative study of E2F transcriptional dynamics at the unmarried-jail cell level past using engineered fluorescent reporter cells provided a quantitative framework for understanding the control logic of prison cell bike entry, challenging the approved textbook model. Genes that regulate the aamplitude of E2F accumulation, such as Myc, decide the delivery in cell cycle and S phase entry. G1 cyclin-CDK activities are not the driver of cell bike entry. Instead, they primarily tune the timing of E2F increment, thereby modulating the footstep of cell bike progression.[fifteen]
Inhibitors [edit]
Endogenous [edit]
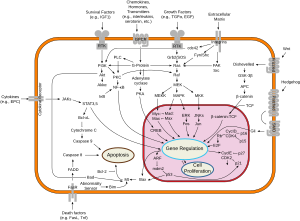
Overview of signal transduction pathways involved in apoptosis, as well known as "programmed cell death"
Two families of genes, the cip/kip (CDK interacting protein/Kinase inhibitory poly peptide) family and the INK4a/ARF (Inhibitor of Kinase 4/Alternative Reading Frame) family unit, foreclose the progression of the jail cell cycle. Considering these genes are instrumental in prevention of tumor germination, they are known as tumor suppressors.
The cip/kip family includes the genes p21, p27 and p57. They halt the cell wheel in Thousandane phase by bounden to and inactivating cyclin-CDK complexes. p21 is activated by p53 (which, in turn, is triggered past Dna impairment eastward.g. due to radiation). p27 is activated past Transforming Growth Factor β (TGF β), a growth inhibitor.
The INK4a/ARF family includes p16INK4a, which binds to CDK4 and arrests the prison cell wheel in G1 stage, and p14ARF which prevents p53 deposition.
Synthetic [edit]
Synthetic inhibitors of Cdc25 could also be useful for the arrest of cell bike and therefore be useful equally antineoplastic and anticancer agents.[28]
Many human cancers possess the hyper-activated Cdk 4/half-dozen activities.[29] Given the observations of cyclin D-Cdk 4/6 functions, inhibition of Cdk iv/6 should outcome in preventing a malignant tumor from proliferating. Consequently, scientists have tried to invent the constructed Cdk4/6 inhibitor every bit Cdk4/6 has been characterized to exist a therapeutic target for anti-tumor effectiveness. Three Cdk4/6 inhibitors - palbociclib, ribociclib, and abemaciclib - currently received FDA approval for clinical use to care for advanced-stage or metastatic, hormone-receptor-positive (Hour-positive, 60 minutes+), HER2-negative (HER2-) breast cancer.[30] [31] For instance, palbociclib is an orally active CDK4/half-dozen inhibitor which has demonstrated improved outcomes for ER-positive/HER2-negative advanced breast cancer. The main side effect is neutropenia which can be managed by dose reduction.[32]
Cdk4/6 targeted therapy volition simply treat cancer types where Rb is expressed. Cancer cells with loss of Rb take primary resistance to Cdk4/6 inhibitors.
Transcriptional regulatory network [edit]
Current show suggests that a semi-autonomous transcriptional network acts in concert with the CDK-cyclin mechanism to regulate the prison cell cycle. Several factor expression studies in Saccharomyces cerevisiae have identified 800–1200 genes that change expression over the class of the cell cycle.[13] [33] [34] They are transcribed at loftier levels at specific points in the cell bicycle, and remain at lower levels throughout the balance of the cycle. While the set of identified genes differs between studies due to the computational methods and criteria used to identify them, each study indicates that a large portion of yeast genes are temporally regulated.[35]
Many periodically expressed genes are driven past transcription factors that are likewise periodically expressed. Ane screen of single-gene knockouts identified 48 transcription factors (about 20% of all non-essential transcription factors) that show cell bike progression defects.[36] Genome-broad studies using loftier throughput technologies have identified the transcription factors that bind to the promoters of yeast genes, and correlating these findings with temporal expression patterns have allowed the identification of transcription factors that drive phase-specific factor expression.[33] [37] The expression profiles of these transcription factors are driven by the transcription factors that elevation in the prior phase, and computational models take shown that a CDK-democratic network of these transcription factors is sufficient to produce steady-land oscillations in gene expression).[34] [38]
Experimental evidence also suggests that gene expression can oscillate with the period seen in dividing wild-type cells independently of the CDK machinery. Orlando et al. used microarrays to measure the expression of a set of 1,271 genes that they identified as periodic in both wild type cells and cells lacking all S-phase and mitotic cyclins (clb1,2,three,iv,5,6). Of the one,271 genes assayed, 882 connected to be expressed in the cyclin-scarce cells at the same time as in the wild type cells, despite the fact that the cyclin-deficient cells arrest at the border between G1 and Southward stage. However, 833 of the genes assayed changed beliefs between the wild blazon and mutant cells, indicating that these genes are probable directly or indirectly regulated past the CDK-cyclin mechanism. Some genes that continued to be expressed on time in the mutant cells were also expressed at different levels in the mutant and wild type cells. These findings suggest that while the transcriptional network may oscillate independently of the CDK-cyclin oscillator, they are coupled in a manner that requires both to ensure the proper timing of cell cycle events.[34] Other piece of work indicates that phosphorylation, a post-translational modification, of cell cycle transcription factors by Cdk1 may alter the localization or activity of the transcription factors in order to tightly control timing of target genes.[36] [39] [40]
While oscillatory transcription plays a key role in the progression of the yeast cell cycle, the CDK-cyclin machinery operates independently in the early embryonic cell cycle. Earlier the midblastula transition, zygotic transcription does non occur and all needed proteins, such as the B-type cyclins, are translated from maternally loaded mRNA.[41]
Deoxyribonucleic acid replication and DNA replication origin activeness [edit]
Analyses of synchronized cultures of Saccharomyces cerevisiae under weather that prevent Deoxyribonucleic acid replication initiation without delaying cell cycle progression showed that origin licensing decreases the expression of genes with origins near their 3' ends, revealing that downstream origins can regulate the expression of upstream genes.[42] This confirms previous predictions from mathematical modeling of a global causal coordination between Dna replication origin activity and mRNA expression,[43] [44] [45] and shows that mathematical modeling of DNA microarray data can exist used to correctly predict previously unknown biological modes of regulation.
Checkpoints [edit]
Prison cell cycle checkpoints are used by the jail cell to monitor and regulate the progress of the cell wheel.[46] Checkpoints forbid cell cycle progression at specific points, allowing verification of necessary phase processes and repair of DNA damage. The cell cannot proceed to the side by side phase until checkpoint requirements take been met. Checkpoints typically consist of a network of regulatory proteins that monitor and dictate the progression of the jail cell through the unlike stages of the cell cycle.
It is estimated that in normal human cells about 1% of single-strand Deoxyribonucleic acid damages are converted to about l endogenous Dna double-strand breaks per cell per cell cycle.[47] Although such double-strand breaks are normally repaired with high fidelity, errors in their repair are considered to contribute significantly to the rate of cancer in humans.[47]
At that place are several checkpoints to ensure that damaged or incomplete Deoxyribonucleic acid is not passed on to girl cells. 3 main checkpoints exist: the K1/S checkpoint, the Gii/Yard checkpoint and the metaphase (mitotic) checkpoint. Another checkpoint is the Go checkpoint, in which the cells are checked for maturity. If the cells fail to laissez passer this checkpoint by not being fix nevertheless, they will be discarded from dividing.
G1/Southward transition is a rate-limiting step in the cell bike and is also known as restriction point.[14] This is where the cell checks whether it has enough raw materials to fully replicate its DNA (nucleotide bases, DNA synthase, chromatin, etc.). An unhealthy or malnourished cell will get stuck at this checkpoint.
The Gtwo/Chiliad checkpoint is where the prison cell ensures that it has enough cytoplasm and phospholipids for ii daughter cells. But sometimes more importantly, information technology checks to meet if it is the right time to replicate. There are some situations where many cells need to all replicate simultaneously (for case, a growing embryo should have a symmetric jail cell distribution until it reaches the mid-blastula transition). This is washed by decision-making the G2/M checkpoint.
The metaphase checkpoint is a fairly minor checkpoint, in that once a cell is in metaphase, it has committed to undergoing mitosis. All the same that'due south not to say it isn't important. In this checkpoint, the cell checks to ensure that the spindle has formed and that all of the chromosomes are aligned at the spindle equator earlier anaphase begins.[48]
While these are the three "primary" checkpoints, not all cells take to pass through each of these checkpoints in this order to replicate. Many types of cancer are caused by mutations that allow the cells to speed through the various checkpoints or fifty-fifty skip them altogether. Going from S to M to S phase almost consecutively. Because these cells have lost their checkpoints, any DNA mutations that may take occurred are disregarded and passed on to the girl cells. This is one reason why cancer cells take a tendency to exponentially accumulate mutations. Aside from cancer cells, many fully differentiated cell types no longer replicate and then they leave the prison cell bicycle and stay in One thousand0 until their death. Thus removing the need for cellular checkpoints. An alternative model of the cell cycle response to Dna impairment has also been proposed, known equally the postreplication checkpoint.
Checkpoint regulation plays an of import role in an organism's development. In sexual reproduction, when egg fertilization occurs, when the sperm binds to the egg, it releases signalling factors that notify the egg that information technology has been fertilized. Among other things, this induces the now fertilized oocyte to return from its previously dormant, G0, state back into the prison cell bicycle and on to mitotic replication and sectionalization.
p53 plays an important role in triggering the control mechanisms at both Thou1/S and Thousand2/M checkpoints. In add-on to p53, checkpoint regulators are existence heavily researched for their roles in cancer growth and proliferation.
Fluorescence imaging of the jail cell wheel [edit]

Fluorescent proteins visualize the cell cycle progression. IFP2.0-hGem(1/110) fluorescence is shown in greenish and highlights the South/G2/Grand phases. smURFP-hCdtI(30/120) fluorescence is shown in carmine and highlights the Thou0/G1 phases.
Pioneering piece of work by Atsushi Miyawaki and coworkers adult the fluorescent ubiquitination-based cell cycle indicator (FUCCI), which enables fluorescence imaging of the prison cell bike. Originally, a green fluorescent protein, mAG, was fused to hGem(one/110) and an orange fluorescent poly peptide (mKO2) was fused to hCdt1(xxx/120). Note, these fusions are fragments that contain a nuclear localization signal and ubiquitination sites for degradation, but are not functional proteins. The green fluorescent protein is made during the S, Grand2, or Grand phase and degraded during the G0 or Thousand1 phase, while the orange fluorescent poly peptide is fabricated during the K0 or One thousand1 phase and destroyed during the S, G2, or K stage.[49] A far-ruby-red and near-infrared FUCCI was developed using a blue-green alga-derived fluorescent protein (smURFP) and a bacteriophytochrome-derived fluorescent protein (movie found at this link).[50]
Role in tumor formation [edit]
A disregulation of the cell cycle components may lead to tumor formation.[51] Every bit mentioned above, when some genes like the cell cycle inhibitors, RB, p53 etc. mutate, they may cause the jail cell to multiply uncontrollably, forming a tumor. Although the duration of cell cycle in tumor cells is equal to or longer than that of normal jail cell cycle, the proportion of cells that are in active cell division (versus quiescent cells in G0 stage) in tumors is much college than that in normal tissue.[52] Thus there is a cyberspace increase in cell number as the number of cells that die past apoptosis or senescence remains the same.
The cells which are actively undergoing cell wheel are targeted in cancer therapy as the DNA is relatively exposed during cell partition and hence susceptible to damage by drugs or radiations. This fact is made utilise of in cancer treatment; by a procedure known every bit debulking, a significant mass of the tumor is removed which pushes a significant number of the remaining tumor cells from One thousand0 to Chiliadi phase (due to increased availability of nutrients, oxygen, growth factors etc.). Radiation or chemotherapy following the debulking procedure kills these cells which take newly entered the cell bike.[14]
The fastest cycling mammalian cells in culture, crypt cells in the intestinal epithelium, have a cycle fourth dimension as short every bit ix to ten hours. Stem cells in resting mouse skin may have a cycle time of more than than 200 hours. Most of this departure is due to the varying length of 1000i, the most variable phase of the wheel. G and S practise not vary much.
In general, cells are most radiosensitive in late Thou and Chiliad2 phases and most resistant in late S phase.
For cells with a longer prison cell cycle time and a significantly long Thou1 stage, there is a 2nd peak of resistance late in Chiliadi.
The design of resistance and sensitivity correlates with the level of sulfhydryl compounds in the jail cell. Sulfhydryls are natural substances that protect cells from radiations damage and tend to be at their highest levels in S and at their everyman near mitosis.
Homologous recombination (HR) is an accurate process for repairing Dna double-strand breaks. HR is nearly absent-minded in G1 phase, is most active in S phase, and declines in G2/Grand.[53] Non-homologous cease joining, a less accurate and more mutagenic procedure for repairing double strand breaks, is active throughout the cell cycle.
See also [edit]
- Cellular model
- Eukaryotic DNA replication
- Origin recognition complex
- Retinoblastoma protein
- Synchronous culture – synchronization of cell cultures
- Wee1
References [edit]
- ^ Wang JD, Levin PA (November 2009). "Metabolism, cell growth and the bacterial jail cell cycle". Nature Reviews. Microbiology. 7 (11): 822–7. doi:10.1038/nrmicro2202. PMC2887316. PMID 19806155.
- ^ Cooper GM (2000). "Chapter 14: The Eukaryotic Cell Wheel". The cell: a molecular approach (2d ed.). Washington, D.C: ASM Press. ISBN978-0-87893-106-iv.
- ^ Smith JA, Martin L (Apr 1973). "Do cells bicycle?". Proceedings of the National Academy of Sciences of the United States of America. seventy (4): 1263–7. Bibcode:1973PNAS...70.1263S. doi:10.1073/pnas.70.four.1263. PMC433472. PMID 4515625.
- ^ Wu RS, Bonner WM (December 1981). "Separation of basal histone synthesis from Due south-phase histone synthesis in dividing cells". Cell. 27 (2 Pt 1): 321–thirty. doi:ten.1016/0092-8674(81)90415-3. PMID 7199388. S2CID 12215040.
- ^ Nelson DM, Ye X, Hall C, Santos H, Ma T, Kao GD, et al. (November 2002). "Coupling of DNA synthesis and histone synthesis in S stage independent of cyclin/cdk2 action". Molecular and Cellular Biology. 22 (21): 7459–72. doi:10.1128/MCB.22.21.7459-7472.2002. PMC135676. PMID 12370293.
- ^ Cameron IL, Greulich RC (July 1963). "Evidence for an essentially constant elapsing of DNA synthesis in renewing epithelia of the adult mouse". The Journal of Cell Biology. 18: 31–xl. doi:ten.1083/jcb.eighteen.1.31. PMC2106275. PMID 14018040.
- ^ Rubenstein, Irwin; Wick, Susan M. (2008). "Jail cell". Earth Book Online Reference Center. Archived from the original on 30 May 2011. Retrieved ten July 2009.
- ^ Maton A, Lahart D, Hopkins J, Warner MQ, Johnson S, Wright JD (1997). Cells: Building Blocks of Life. New Jersey: Prentice Hall. pp. 70–4. ISBN978-0-13-423476-2.
- ^ De Souza CP, Osmani SA (September 2007). "Mitosis, non only open or closed". Eukaryotic Cell. 6 (9): 1521–7. doi:10.1128/EC.00178-07. PMC2043359. PMID 17660363.
- ^ Lilly MA, Duronio RJ (April 2005). "New insights into cell cycle command from the Drosophila endocycle". Oncogene. 24 (17): 2765–75. doi:10.1038/sj.onc.1208610. PMID 15838513.
- ^ Nigg EA (June 1995). "Cyclin-dependent protein kinases: key regulators of the eukaryotic jail cell cycle". BioEssays. 17 (6): 471–80. doi:x.1002/bies.950170603. PMID 7575488. S2CID 44307473.
- ^ "Press release". Nobelprize.org.
- ^ a b Spellman PT, Sherlock G, Zhang MQ, Iyer VR, Anders Yard, Eisen MB, et al. (December 1998). "Comprehensive identification of prison cell cycle-regulated genes of the yeast Saccharomyces cerevisiae by microarray hybridization". Molecular Biology of the Cell. 9 (12): 3273–97. doi:10.1091/mbc.9.12.3273. PMC25624. PMID 9843569.
- ^ a b c d Robbins SL, Cotran RS (2004). Kumar V, Abbas AK, Fausto N (eds.). Pathological Basis of Disease. Elsevier. ISBN978-81-8147-528-2.
- ^ a b Dong P, Maddali MV, Srimani JK, Thélot F, Nevins JR, Mathey-Prevot B, You lot L (September 2014). "Division of labour betwixt Myc and G1 cyclins in cell cycle delivery and stride control". Nature Communications. v: 4750. Bibcode:2014NatCo...5.4750D. doi:10.1038/ncomms5750. PMC4164785. PMID 25175461.
- ^ Mahmoudi M, Azadmanesh One thousand, Shokrgozar MA, Journeay WS, Laurent S (May 2011). "Event of nanoparticles on the cell life cycle". Chemical Reviews. 111 (5): 3407–32. doi:10.1021/cr1003166. PMID 21401073.
- ^ Goel South, DeCristo MJ, McAllister SS, Zhao JJ (November 2018). "CDK4/6 Inhibition in Cancer: Beyond Cell Cycle Arrest". Trends in Prison cell Biological science. 28 (11): 911–925. doi:x.1016/j.tcb.2018.07.002. PMC6689321. PMID 30061045.
- ^ Burkhart DL, Sage J (September 2008). "Cellular mechanisms of neoplasm suppression by the retinoblastoma gene". Nature Reviews. Cancer. eight (9): 671–82. doi:10.1038/nrc2399. PMC6996492. PMID 18650841.
- ^ Morgan Practise (2007). The cell bicycle : principles of control. London: New Science Press. ISBN978-0-xix-920610-0. OCLC 70173205.
- ^ Paternot S, Bockstaele Fifty, Bisteau Ten, Kooken H, Coulonval One thousand, Roger PP (February 2010). "Rb inactivation in cell cycle and cancer: the puzzle of highly regulated activating phosphorylation of CDK4 versus constitutively active CDK-activating kinase". Cell Wheel. 9 (4): 689–99. doi:10.4161/cc.nine.four.10611. PMID 20107323.
- ^ Henley SA, Dick FA (March 2012). "The retinoblastoma family of proteins and their regulatory functions in the mammalian cell division cycle". Jail cell Partitioning. 7 (i): ten. doi:10.1186/1747-1028-7-10. PMC3325851. PMID 22417103.
- ^ a b c Narasimha AM, Kaulich M, Shapiro GS, Choi YJ, Sicinski P, Dowdy SF (June 2014). "Cyclin D activates the Rb tumor suppressor by mono-phosphorylation". eLife. 3: e02872. doi:10.7554/eLife.02872. PMC4076869. PMID 24876129.
- ^ Morris EJ, Dyson NJ (1 January 2001). Retinoblastoma poly peptide partners. Advances in Cancer Inquiry. Vol. 82. Academic Press. pp. one–54. doi:ten.1016/s0065-230x(01)82001-7. ISBN9780120066827. PMID 11447760.
- ^ Dyson NJ (July 2016). "RB1: a prototype tumor suppressor and an enigma". Genes & Development. thirty (13): 1492–502. doi:ten.1101/gad.282145.116. PMC4949322. PMID 27401552.
- ^ a b Sanidas I, Morris R, Fella KA, Rumde PH, Boukhali Thousand, Tai EC, et al. (March 2019). "A Code of Mono-phosphorylation Modulates the Function of RB". Molecular Cell. 73 (5): 985–1000.e6. doi:10.1016/j.molcel.2019.01.004. PMC6424368. PMID 30711375.
- ^ a b Topacio BR, Zatulovskiy Eastward, Cristea S, Xie Southward, Tambo CS, Rubin SM, et al. (May 2019). "Cyclin D-Cdk4,6 Drives Jail cell-Bicycle Progression via the Retinoblastoma Protein'southward C-Last Helix". Molecular Jail cell. 74 (iv): 758–770.e4. doi:ten.1016/j.molcel.2019.03.020. PMC6800134. PMID 30982746.
- ^ Norbury C (1995). "Cdk2 protein kinase (vertebrates)". In Hardie DG, Hanks S (eds.). Protein kinase factsBook. Boston: Academic Press. pp. 184. ISBN978-0-12-324719-three.
- ^ "Presentation on CDC25 PHOSPHATASES: A Potential Target for Novel Anticancer Agents". Archived from the original on iii March 2016. Retrieved 11 March 2010.
- ^ Sherr CJ, Beach D, Shapiro GI (Apr 2016). "Targeting CDK4 and CDK6: From Discovery to Therapy". Cancer Discovery. 6 (4): 353–67. doi:x.1158/2159-8290.cd-fifteen-0894. PMC4821753. PMID 26658964.
- ^ O'Leary B, Finn RS, Turner NC (July 2016). "Treating cancer with selective CDK4/six inhibitors". Nature Reviews. Clinical Oncology. thirteen (7): 417–30. doi:x.1038/nrclinonc.2016.26. PMID 27030077. S2CID 23646632.
- ^ Bilgin B, Sendur MA, Şener Dede D, Akıncı MB, Yalçın B (September 2017). "A current and comprehensive review of cyclin-dependent kinase inhibitors for the treatment of metastatic breast cancer". Current Medical Research and Opinion. 33 (9): 1559–1569. doi:10.1080/03007995.2017.1348344. PMID 28657360. S2CID 205542255.
- ^ Schmidt Thou, Sebastian M (August 2018). "Palbociclib-The First of a New Class of Jail cell Cycle Inhibitors". Recent Results in Cancer Research. Fortschritte der Krebsforschung. Progres dans les Recherches Sur le Cancer. Recent Results in Cancer Research. 211: 153–175. doi:10.1007/978-3-319-91442-8_11. ISBN978-3-319-91441-1. PMID 30069766.
- ^ a b Pramila T, Wu W, Miles S, Noble WS, Breeden LL (August 2006). "The Forkhead transcription factor Hcm1 regulates chromosome segregation genes and fills the S-phase gap in the transcriptional circuitry of the cell wheel". Genes & Evolution. 20 (16): 2266–78. doi:10.1101/gad.1450606. PMC1553209. PMID 16912276.
- ^ a b c Orlando DA, Lin CY, Bernard A, Wang JY, Socolar JE, Iversen ES, et al. (June 2008). "Global control of prison cell-bike transcription by coupled CDK and network oscillators". Nature. 453 (7197): 944–seven. Bibcode:2008Natur.453..944O. doi:x.1038/nature06955. PMC2736871. PMID 18463633.
- ^ de Lichtenberg U, Jensen LJ, Fausbøll A, Jensen TS, Bork P, Brunak Due south (April 2005). "Comparing of computational methods for the identification of jail cell wheel-regulated genes". Bioinformatics. 21 (7): 1164–71. doi:ten.1093/bioinformatics/bti093. PMID 15513999.
- ^ a b White MA, Riles L, Cohen BA (February 2009). "A systematic screen for transcriptional regulators of the yeast cell wheel". Genetics. 181 (2): 435–46. doi:10.1534/genetics.108.098145. PMC2644938. PMID 19033152.
- ^ Lee TI, Rinaldi NJ, Robert F, Odom DT, Bar-Joseph Z, Gerber GK, et al. (October 2002). "Transcriptional regulatory networks in Saccharomyces cerevisiae". Science. 298 (5594): 799–804. Bibcode:2002Sci...298..799L. doi:10.1126/science.1075090. PMID 12399584. S2CID 4841222.
- ^ Simon I, Barnett J, Hannett N, Harbison CT, Rinaldi NJ, Volkert TL, et al. (September 2001). "Series regulation of transcriptional regulators in the yeast cell bicycle". Jail cell. 106 (vi): 697–708. doi:10.1016/S0092-8674(01)00494-9. PMID 11572776. S2CID 9308235.
- ^ Sidorova JM, Mikesell GE, Breeden LL (December 1995). "Prison cell bicycle-regulated phosphorylation of Swi6 controls its nuclear localization". Molecular Biology of the Prison cell. 6 (12): 1641–58. doi:10.1091/mbc.6.12.1641. PMC301322. PMID 8590795.
- ^ Ubersax JA, Woodbury EL, Quang PN, Paraz M, Blethrow JD, Shah Yard, et al. (October 2003). "Targets of the cyclin-dependent kinase Cdk1". Nature. 425 (6960): 859–64. Bibcode:2003Natur.425..859U. doi:10.1038/nature02062. PMID 14574415. S2CID 4391711.
- ^ Morgan DO (2007). "ii–3". The Prison cell Cycle: Principles of Command. London: New Science Printing. p. 18. ISBN978-0-9539181-two-6.
- ^ Omberg Fifty, Meyerson JR, Kobayashi Thou, Drury LS, Diffley JF, Alter O (October 2009). "Global furnishings of DNA replication and DNA replication origin activity on eukaryotic gene expression". Molecular Systems Biological science. v: 312. doi:ten.1038/msb.2009.seventy. PMC2779084. PMID 19888207.
- ^ Alter O, Golub GH, Chocolate-brown PO, Botstein D (2004). Deutscher MP, Black South, Boehmer PE, D'Urso G, Fletcher TM, Huijing F, Marshall A, Pulverer B, Renault B, Rosenblatt JD, Slingerland JM, Whelan WJ (eds.). "Novel Genome-Scale Correlation betwixt Dna Replication and RNA Transcription During the Cell Bicycle in Yeast is Predicted by Data-Driven Models" (PDF). Miami Nature Biotechnology Winter Symposium: Cell Cycle, Chromosomes and Cancer. Miami Embankment, FL: University of Miami School of Medicine, vol. xv (Jan 31 – Feb 4, 2004).
- ^ Alter O, Golub GH (November 2004). "Integrative assay of genome-scale data by using pseudoinverse projection predicts novel correlation between DNA replication and RNA transcription". Proceedings of the National Academy of Sciences of the United States of America. 101 (47): 16577–82. Bibcode:2004PNAS..10116577A. doi:ten.1073/pnas.0406767101. PMC534520. PMID 15545604.
- ^ Omberg Fifty, Golub GH, Alter O (Nov 2007). "A tensor higher-order singular value decomposition for integrative assay of DNA microarray data from different studies". Proceedings of the National Academy of Sciences of the United States of America. 104 (47): 18371–half dozen. Bibcode:2007PNAS..10418371O. doi:10.1073/pnas.0709146104. PMC2147680. PMID 18003902.
- ^ Elledge SJ (December 1996). "Prison cell bike checkpoints: preventing an identity crisis". Scientific discipline. 274 (5293): 1664–72. Bibcode:1996Sci...274.1664E. doi:x.1126/science.274.5293.1664. PMID 8939848. S2CID 39235426.
- ^ a b Vilenchik MM, Knudson AG. Endogenous Dna double-strand breaks: production, fidelity of repair, and induction of cancer. Proc Natl Acad Sci U Due south A. 2003 Oct 28;100(22):12871-6. doi: 10.1073/pnas.2135498100. Epub 2003 October 17. PMID 14566050; PMCID: PMC240711.
- ^ LeMaire-Adkins R, Radke K, Hunt PA (December 1997). "Lack of checkpoint control at the metaphase/anaphase transition: a mechanism of meiotic nondisjunction in mammalian females". The Journal of Cell Biology. 139 (7): 1611–nine. doi:10.1083/jcb.139.seven.1611. PMC2132649. PMID 9412457.
- ^ Sakaue-Sawano A, Kurokawa H, Morimura T, Hanyu A, Hama H, Osawa H, et al. (February 2008). "Visualizing spatiotemporal dynamics of multicellular cell-wheel progression". Cell. 132 (iii): 487–98. doi:10.1016/j.cell.2007.12.033. PMID 18267078. S2CID 15704902.
- ^ Rodriguez EA, Tran GN, Gross LA, Crisp JL, Shu X, Lin JY, Tsien RY (September 2016). "A far-red fluorescent poly peptide evolved from a cyanobacterial phycobiliprotein". Nature Methods. 13 (9): 763–9. doi:10.1038/nmeth.3935. PMC5007177. PMID 27479328.
- ^ Champeris Tsaniras S, Kanellakis N, Symeonidou IE, Nikolopoulou P, Lygerou Z, Taraviras S (June 2014). "Licensing of DNA replication, cancer, pluripotency and differentiation: an interlinked world?". Seminars in Jail cell & Developmental Biology. xxx: 174–eighty. doi:x.1016/j.semcdb.2014.03.013. PMID 24641889.
- ^ Baserga, Renaro (June 1965). "The Relationship of the Cell Cycle to Tumor Growth and Command of Prison cell Division". Cancer Research. 25: 581–595.
- ^ Mao Z, Bozzella K, Seluanov A, Gorbunova V (September 2008). "DNA repair by nonhomologous terminate joining and homologous recombination during cell bike in human cells". Jail cell Cycle. 7 (18): 2902–6. doi:x.4161/cc.7.18.6679. PMC2754209. PMID 18769152.
Farther reading [edit]
- Morgan DO (2007). The Cell Cycle: Principles of Control. London: Published by New Scientific discipline Press in association with Oxford University Press. ISBN978-0-87893-508-6.
- Alberts B, Johnson A, Lewis J, Raff Yard, Roberts G, Walter P (2008). "Chapter 17". Molecular Biology of the Prison cell (fifth ed.). New York: Garland Science. ISBN978-0-8153-4111-6.
- Krieger 1000, Scott MP, Matsudaira PT, Lodish HF, Darnell JE, Zipursky L, Kaiser C, Berk A (2004). Molecular cell biology. New York: Due west.H. Freeman and CO. ISBN978-0-7167-4366-8.
- Watson JD, Baker TA, Bell SP, Gann A, Levine M, Losick R (2004). "Chapter 7". Molecular biology of the gene (5th ed.). San Francisco: Pearson/Benjamin Cummings. ISBN978-0-8053-4642-8.
External links [edit]
| | Wikimedia Commons has media related to Prison cell cycle. |
-
 This article incorporates public domain cloth from the NCBI certificate: "Science Primer".
This article incorporates public domain cloth from the NCBI certificate: "Science Primer". - David Morgan'due south Seminar: Decision-making the Cell Wheel
- The cell cycle & Cell death
- Transcriptional program of the cell cycle: high-resolution timing
- Cell cycle and metabolic cycle regulated transcription in yeast
- Cell Cycle Animation 1Lec.com
- Cell Cycle
- Fucci:Using GFP to visualize the cell-bike
- Science Creative Quarterly'south overview of the prison cell cycle
- KEGG – Man Cell Cycle
Source: https://en.wikipedia.org/wiki/Cell_cycle
0 Response to "Which Stage Is Most Associated With a Cell That Is Unable to Divide Again Such as a Nerve Cell"
Post a Comment